HTRA2 and its role in parkinson’s disease: genetic and functional insights
HTRA2 and its role in parkinson’s disease
Authors
Abstract
Aim This study aimed to investigate the genetic variations, protein-protein interactions, and functional involvement of HTRA2 in mitochondrial homeostasis and apoptosis, with a focus on its potential contribution to Parkinson’s disease (PD) pathology.
Methods Genetic variations in HTRA2 were identified using dbSNP, SWISS-Prot, and ClinVar. Protein-protein interaction (PPI) analysis was performed using STRING and GeneMANIA. Gene expression data were obtained from UniProt, BioXpress, and GTEx, and pathway analysis was conducted using KEGG. Disease associations were identified through the DISEASES database.
Results HTRA2 was found to interact strongly with apoptosis-related proteins, including DIABLO, XIAP, BIRC2, and BIRC3. It exhibited high expression in brain regions crucial for neuronal survival. Nine missense mutations were identified as potentially pathogenic. KEGG pathway analysis linked HTRA2 to mitochondrial dysfunction and apoptosis. Additionally, associations with Alzheimer’s disease, essential tremor, and cancer were observed.
Conclusion HTRA2 plays a critical role in maintaining mitochondrial quality control and regulating apoptosis. Its dysfunction may contribute to PD pathogenesis by promoting neuronal death. These findings suggest that HTRA2 may represent a promising therapeutic target for PD and related neurodegenerative disorders.
Keywords
Introduction
HTRA2 (High Temperature Requirement A2) is a serine protease that resides in the intermembrane space of mitochondria, where it plays a crucial role in maintaining mitochondrial homeostasis and regulating apoptotic pathways. This protein is essential for preserving mitochondrial integrity by preventing the accumulation of misfolded or damaged proteins, thereby safeguarding cellular energy production and overall cellular health.1
The HTRA2 gene is located on chromosome 2p13.1 and encodes a 458-amino acid protein that is structurally defined by three key features: an N-terminal Ala-Val-Pro-Ser (AVPS) motif, which is crucial for its mitochondrial localization and activation; a central serine protease domain responsible for its catalytic activity; and a C-terminal PDZ domain, which regulates substrate binding and enzymatic specificity.2 The functional activity of HTRA2 is dependent on its homotrimeric structure, which ensures its stability and optimal enzymatic function.3 HTRA2 serves a dual role as both a mitochondrial quality control protease and a pro-apoptotic factor. Under physiological conditions, it functions as a surveillance system by degrading defective proteins and preventing mitochondrial dysfunction. However, in response to cellular stress, HTRA2 is released into the cytosol, where it promotes apoptosis by binding to inhibitors of apoptosis proteins (IAPs) and facilitating caspase activation. Dysregulation of this finely tuned balance has been implicated in the pathogenesis of various neurodegenerative disorders, most notably Parkinson’s disease (PD) and essential tremor, where mitochondrial dysfunction and defective proteostasis contribute to disease progression.4
Given the critical role of HTRA2 in mitochondrial function and apoptosis, this study aims to explore its potential association with Parkinson’s disease through in silico approaches. Specifically, we will investigate the mitochondrial localization of HTRA2, identify genes with functionally similar roles, and analyze genetic variations within the HTRA2 gene that may contribute to PD pathology. By integrating bioinformatics tools and genetic analysis, this research seeks to enhance our understanding of HTRA2’s involvement in neurodegeneration and its potential as a therapeutic target in Parkinson’s disease.1
Materials and Methods
Finding the HTRA2 Gene Dataset CollectionInformation on the HTRA2 gene was obtained from the Online Mendelian Inheritance in Man (OMIM) and National Center for Biological Information (NCBI) Entrez Gene databases. Data on Single Nucleotide Polymorphisms (SNPs) 5 associated with the HTRA2 gene, including protein accession numbers and SNP IDs, were retrieved from the NCBI dbSNP and SWISS-Prot databases.5
Identification of HTRA2-Related Genes Using the STRING Database
The STRING database integrates protein-protein interaction data from various organisms to provide a comprehensive map of biological networks.6 It offers biologically relevant and specific interaction data, enabling researchers to understand gene functions within cellular pathways. In this study, we utilized the STRING database to identify genes that are biologically, functionally, and physically linked to HTRA2. The findings obtained from STRING were further validated using the GeneMANIA program to ensure consistency and accuracy.7
Investigation of HTRA2 Gene Expression Levels
In this study, SNPs predicted to have damaging effects on HTRA2 were identified, and their allele frequencies were calculated using this database. The SIFT algorithm was employed to determine nonsynonymous variants, while PolyPhen-2 was used to assess their potential damaging effects.8 All results were cross-validated to ensure reliability. Given the significance of gene expression analysis in establishing links between genes and diseases, the expression level of HTRA2 was retrieved from UniProt and BioXpress databases.2 Further verification of gene expression, exon expression, and junction expression was conducted using the GTEx portal and the Ensembl Genome Browser to ensure consistency.
Identification of Mutations in the HTRA2 GeneFor genetic variation analysis, Single Nucleotide Polymorphism (SNP) data related to HTRA2, including protein accession numbers and SNP identifiers, were acquired from NCBI dbSNP and SWISS-Prot.9 Additionally, population allele frequencies and functional consequences of identified variants were assessed using the Genome Aggregation Database (gnomAD) 6 and dbSNP. The pathogenicity and clinical significance of specific variants were further examined using ClinVar2, while potential functional impacts were predicted through Ensembl Variant Effect Predictor (VEP).6
Pathway and Diseases Analysis of HTRA2The Kyoto Encyclopedia of Genes and Genomes (KEGG) database was utilized to systematically analyze the functional pathways associated with the HTRA2 gene. KEGG was employed to map the biological pathways involving HTRA2, highlighting its crucial role in mitochondrial quality control and apoptosis regulation. This database integrates genomic data with other functionally classified biological processes, providing insights into HTRA2’s involvement in cellular homeostasis and disease mechanisms.10
To validate our findings, pathway comparisons were conducted using STRING (Search Tool for the Retrieval of Interacting Genes/ Proteins), which allowed us to identify and confirm protein-protein interactions associated with HTRA2. Furthermore, disease associations related to HTRA2 were investigated using the DISEASES resource database, enabling us to establish links between HTRA2 and neurodegenerative disorders, particularly Parkinson’s disease.4
Ethical ApprovalNot applicable. This study was conducted entirely using publicly available databases and bioinformatics tools, and did not involve human participants, animals, or identifiable personal data.
Statistical Analysis No conventional statistical tests involving human or clinical datasets were performed. Computational predictions andvariant pathogenicity assessments were conducted using established bioinformatics algorithms, including SIFT and PolyPhen-2. Gene expression and interaction analyses were performed using publicly available databases and validated through cross-database comparisons.
Reporting GuidelinesThis study is reported in accordance with the STROBE guideline.
Results
Identification of HTRA2-Related Genes Using the STRING DatabaseSTRING analysis revealed strong associations between HTRA2 and apoptosis-related proteins, particularly XIAP, BIRC2, and BIRC3, confirming its role in apoptotic regulation. Figure 1A shows that the protein-protein interaction (PPI) network analysis revealed a highly interconnected network of apoptosis-related proteins, underscoring key regulatory molecules such as DIABLO (SMAC), XIAP, BIRC2, BIRC3, and BIRC6. The strong associations observed among these proteins suggest their pivotal roles in apoptotic signaling pathways. Notably, DIABLO and HTRA2 exhibited multiple interactions with inhibitors of apoptosis proteins (IAPs), indicating their involvement in the promotion of apoptosis through the antagonism of XIAP and related proteins. Furthermore, PINK1 and PARL, primarily known for their roles in mitochondrial quality control, demonstrated significant interactions with apoptotic regulators, suggesting a potential mechanistic link between mitochondrial dysfunction and apoptosis. The presence of TNFRSF1A, a key component of the tumor necrosis factor (TNF) signaling pathway, further highlights the interplay between extrinsic and intrinsic apoptotic mechanisms. The dense connectivity within the network underscores the intricate regulatory crosstalk among these proteins, reinforcing their functional significance in apoptosis regulation and mitochondrial homeostasis.
Investigation of HTRA2 Gene Expression LevelsHTRA2 exhibited high expression in the cerebellum, cerebral cortex, and hippocampus, supporting its involvement in neuronal survival. The analysis of HTRA2 protein expression across various tissues revealed a widespread but variable distribution, with notable differences among organ systems (Figure 1B).
Figure 1B shows that the high to moderate expression levels were observed in the central nervous system, particularly in the cerebellum, cerebral cortex, and hippocampus, suggesting a potential role in neuronal function and homeostasis. Additionally, significant expressions were detected in the gastrointestinal tract, pancreas, kidneys, and reproductive organs, indicating their involvement in metabolic and apoptotic processes in these tissues. In contrast, lower expression levels were noted in skeletal muscle, soft tissue, and lymphoid organs, while minimal or undetectable expression was observed in certain tissues such as the testis and specific immune-related structures. Furthermore, clustering analysis placed HTRA2 within a gene group associated with fundamental cellular processes, reinforcing its functional relevance in apoptosis and mitochondrial quality control. These findings support the hypothesis that HTRA2 plays a critical role in mitochondrial homeostasis and apoptotic regulation, particularly in metabolically active and neuroprotective environments.
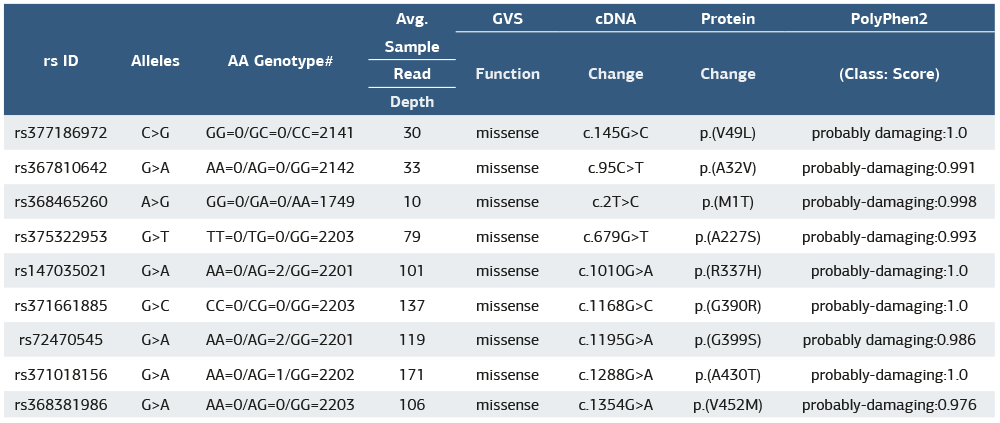
Nine missense mutations were identified as potentially damaging, with high PolyPhen2 scores suggesting pathogenic effects. These mutations may impair HTRA2’s protease activity, affecting mitochondrial function and apoptosis regulation. Table 1 presents a list of identified missense mutations in the HTRA2 gene, including their corresponding reference SNP (rs) IDs, nucleotide variations (alleles), amino acid genotypes, sequencing read depth, cDNA and protein changes, and predicted functional impact based on PolyPhen2 scores. Each mutation results in an amino acid substitution, which may affect HTRA2’s function in mitochondrial quality control and apoptosis regulation. The PolyPhen2 scores, ranging close to 1.0, indicate that these mutations are likely to be damaging, suggesting a potential role in disease mechanisms. This data highlights the importance of these variants in understanding HTRA2-related dysfunctions, particularly in neurodegenerative conditions such as Parkinson’s disease.
Pathway and Diseases Analysis of HTRA2KEGG analysis confirmed HTRA2’s role in mitochondrial dysfunction and apoptosis, while the DISEASES database linked HTRA2 to PD, Alzheimer’s disease, essential tremor, and cancer, suggesting broader implications beyond PD. HTRA2 was identified as a key player in the mitochondrial dysfunction and apoptosis pathways associated with Parkinson’s disease (PD). The pathway analysis revealed that HTRA2 interacts with PINK1 and PARK2, two crucial regulators of mitochondrial quality control, indicating its role in mitophagy and proteostasis. Under normal conditions, HTRA2 functions as a mitochondrial serine protease, preventing the accumulation of misfolded proteins. However, in the context of PD, HTRA2 dysfunction contributes to increased oxidative stress, ATP depletion, and neuronal apoptosis. Additionally, HTRA2 is released into the cytosol upon mitochondrial stress, where it promotes caspase activation and cell death. These findings highlight the dual role of HTRA2 in mitochondrial maintenance and programmed cell death, suggesting that its dysregulation may accelerate dopaminergic neuron degeneration in Parkinson’s disease. The association between HTRA2, PINK1, and PARK2 mutations further supports its pathogenic significance in PD, making it a potential therapeutic target for neuroprotection (Figure 2).
The DISEASES database analysis highlights a strong association between HTRA2 and multiple pathological conditions, particularly neurodegenerative diseases (Figure 3). The highest Z-score and confidence rating were observed for Parkinson’s disease (PD) and Parkinsonism, reinforcing HTRA2’s crucial role in mitochondrial dysfunction, apoptosis, and protein quality control, which are key mechanisms in PD pathology. Additionally, HTRA2 was linked to other neurological disorders, including essential tremor, Alzheimer’s disease, dementia, and neuropathy, suggesting its broader involvement in neuronal maintenance and survival. Beyond neurodegeneration, HTRA2 also showed a significant association with cancer, indicating a possible role in tumor suppression and apoptotic regulation. Its connections to hematopoietic and lymphatic system diseases further suggest potential involvement in immune cell homeostasis and programmed cell death. These findings collectively emphasize HTRA2’s multifaceted role in both neurodegenerative and systemic diseases, making it a potential therapeutic target for future research.
Discussion
The findings from this study highlight the significant role of HTRA2 in Parkinson’s disease (PD) and its potential contribution to neurodegenerative mechanisms. HTRA2 is a mitochondrial serine protease that is released into the cytosol following cellular stress, where it interacts with inhibitor of apoptosis proteins (IAPs) and promotes caspase activation, leading to programmed cell death.11 The STRING-based protein-protein interaction (PPI) analysis identified HTRA2 as a key regulator of apoptosis, interacting with DIABLO (SMAC), XIAP, BIRC2, BIRC3, and BIRC6, reinforcing its role in mitochondrial homeostasis and apoptotic signaling pathways.12 Additionally, HTRA2 was found to interact with PINK1 and PARK2, two crucial regulators of mitophagy, suggesting that HTRA2 dysfunction may exacerbate mitochondrial impairment and neuronal degeneration in PD pathology.13,14
The gene expression analysis demonstrated high to moderate expression levels of HTRA2 in the central nervous system, particularly in the cerebellum, cerebral cortex, and hippocampus, which aligns with its potential role in neuronal maintenance and homeostasis.3 Expression was also observed in other metabolically active tissues, including the pancreas, kidneys, and reproductive organs, indicating its broader function beyond the nervous system. The expression profile of HTRA2 in multiple brain regions suggests its involvement in neuroprotective mechanisms, which may become compromised in neurodegenerative diseases.
Mutational analysis revealed several missense mutations in the HTRA2 gene, with high PolyPhen2 scores indicating potentially damaging effects. Among the identified variants, p.(V49L), p.(A32V), p.(M1T), p.(A227S), and p.(R337H) showed the strongest pathogenic predictions, suggesting their possible involvement in PD susceptibility. Previous studies have reported that mutations in HTRA2 (such as p.G399S and Pro143Ala) are associated with familial and sporadic cases of Parkinson’s disease.15,16,17 Our study expands on this by identifying additional potentially deleterious mutations, reinforcing the genetic contribution of HTRA2 to PD pathogenesis.
Pathway enrichment analysis using KEGG confirmed the involvement of HTRA2 in apoptosis and mitochondrial dysfunction, with PINK1 and PARK2 playing key roles in regulating its function. The DISEASES database further supported the strong correlation between HTRA2 and Parkinson’s disease, with a high Z-score and confidence level. Interestingly, HTRA2 was also linked to other neurological conditions, including Alzheimer’s disease, dementia, and essential tremors, suggesting that its dysfunction may contribute to broader neurodegenerative processes.
Overall, our findings indicate that HTRA2 plays a dual role in mitochondrial quality control and apoptosis, and its dysregulation may contribute to dopaminergic neuron loss in PD. The strong association between HTRA2, PINK1, and PARK2 mutations supports their pathogenic significance in PD and highlights their potential as a therapeutic target for neuroprotection. Future experimental studies are required to validate these findings in cellular and animal models, further elucidating the molecular mechanisms underlying HTRA2 dysfunction in neurodegeneration.
Limitations
This study has several limitations that should be acknowledged. First, the research was based entirely on in silico analyses, including database-derived gene expression profiles, protein– protein interaction networks, and computational predictions of variant pathogenicity. Although such approaches provide valuable insights, they cannot replace functional validation through cellular or animal experiments. Second, the accuracy of variant effect predictions relies heavily on the databases used (dbSNP, ClinVar², gnomAD, STRING, KEGG), all of which are subject to updates, incomplete annotations, and population- specific biases. Third, the study did not incorporate experimental confirmation of protease activity, mitochondrial dysfunction, or apoptosis modulation, limiting the ability to draw definitive conclusions regarding HTRA2’s mechanistic role in Parkinson’s disease. Additionally, gene expression data retrieved from publicly available repositories may not fully capture cell-type– specific expression differences, especially within vulnerable dopaminergic neuronal populations.
Conclusion
HTRA2 plays a critical role in mitochondrial homeostasis and apoptosis regulation, and its dysregulation may contribute to PD pathology. The strong genetic and functional associations between HTRA2, PINK1, and PARK2 reinforce their importance in mitochondrial dysfunction and neurodegeneration. These findings suggest that HTRA2 represents a promising target for therapeutic intervention in PD. Future studies should focus on validating these findings through experimental approaches, exploring HTRA2 inhibitors or activators as potential neuroprotective agents.
Declarations
Ethics Declarations
The authors confirm that this study was performed using publicly available datasets and bioinformatics tools. No human participants, animals, or identifiable personal data were involved.
Animal and Human Rights Statement
Not applicable. This study did not involve human participants or animals.
Informed Consent
Not applicable. The study used publicly available databases and did not involve human participants.
Data Availability
All data analyzed in this study are publicly available from the referenced databases, including NCBI dbSNP, SWISS-Prot, ClinVar, STRING, GeneMANIA, UniProt, BioXpress, GTEx, KEGG, and the DISEASES database.
Conflict of Interest
The authors declare that there is no conflict of interest.
Funding
None.
Author Contributions (CRediT Taxonomy)
Conceptualization: D.K.
Methodology: D.K., A.U.T.
Investigation: D.K., S.K.T.
Data curation: D.K.
Formal analysis: A.U.T.
Writing – original draft: D.K.
Writing – review & editing: D.K., S.K.T.
Supervision: S.K.T.
Scientific Responsibility Statement
The authors declare that they are responsible for the article’s scientific content, including study design, data collection, analysis and interpretation, writing, and some of the main line, or all of the preparation and scientific review of the contents, and approval of the final version of the article.
Abbreviations
AVPS: Ala-Val-Pro-Ser
DIABLO: Direct IAP Binding Protein with Low pI
GTEx: Genotype-Tissue Expression
HTRA2: High Temperature Requirement A2
IAP: Inhibitor of Apoptosis Protein
KEGG: Kyoto Encyclopedia of Genes and Genomes
NCBI: National Center for Biotechnology Information
OMIM: Online Mendelian Inheritance in Man
PD: Parkinson’s Disease
PPI: Protein-Protein Interaction
SNP: Single Nucleotide Polymorphism
STRING: Search Tool for the Retri
References
-
Goo HG, Rhim H, Kang S. HTRA2/Omi influences the stability of LON protease 1 and prohibitin, proteins involved in mitochondrial homeostasis. Exp Cell Res. 2014;328(2):456-465.
-
Kang S, Fernandes-Alnemri T, Alnemri ES. A novel role for the mitochondrial HTRA2/OMI protease in aging. Autophagy. 2013;9(3):420-421.
-
Su XJ, Huang L, Qu Y, Mu D. Progress in research on the role of Omi/HTRA2 in neurological diseases. Rev Neurosci. 2019;30(3):279-287.
-
Nam MK, Seong YM, Park HJ, et al. The homotrimeric structure of HTRA2 is indispensable for executing its serine protease activity. Exp Mol Med. 2006;38(1):36-43. doi:10.1038/emm.2006.5
-
Unal Gulsuner H, Gulsuner S, Mercan FN, et al. Mitochondrial serine protease HTRA2 p.G399S in a kindred with essential tremor and Parkinson’s disease. Proc Natl Acad Sci U S A. 2014;111(51):18285-18290. doi:10.1073/pnas.1419581111
-
Strauss KM, Martins LM, Plun-Favreau H, et al. Loss of function mutations in the gene encoding Omi/HTRA2 in Parkinson’s disease. Hum Mol Genet. 2005;14(15):2099-2111. doi:10.1093/hmg/ddi215
-
Bogaerts V, Nuytemans K, Reumers J, et al. Genetic variability in the mitochondrial serine protease HTRA2 contributes to risk for Parkinson’s disease. Hum Mutat. 2008;29(6):832-840.
-
Szklarczyk D, Morris JH, Cook H, et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45(D1):D362-D368. doi:10.1093/nar/gkw937
-
Warde-Farley D, Donaldson SL, Comes O, et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010;38(Web Server issue):W214-W220. doi:10.1093/nar/gkq537
-
Robinson JT, Thorvaldsdóttir H, Winckler W, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29(1):24-26. doi:10.1038/nbt.1754
-
Verhagen AM, Silke J, Ekert PG, et al. HtrA2 promotes cell death through its serine protease activity and its ability to antagonize inhibitor of apoptosis proteins. J Biol Chem. 2002;277(1):445-454.
-
Liu Z, Sun C, Olejniczak ET, et al. Structural basis for binding of Smac/DIABLO to the XIAP BIR3 domain. Nature. 2000;408(6815):1004-1008. doi:10.1038/35050006
-
Liu J, Liu W, Li R, Yang H. Mitophagy in Parkinson’s disease: from pathogenesis to treatment. Cells. 2019;8(7):712. doi:10.3390/cells8070712
-
Zong Y, Li H, Liao P, et al. Mitochondrial dysfunction: mechanisms and advances in therapy. Signal Transduct Target Ther. 2024;9(1):124.
-
Darreh-Shori T, Rezaeianyazdi S, Lana E, et al. Increased active OMI/HTRA2 serine protease displays a positive correlation with cholinergic alterations in the Alzheimer’s disease brain. Mol Neurobiol. 2019;56(7):4601-4619.
-
Bose K, Wagh A, Mishra V, et al. Loss of GSK-3β-mediated phosphorylation in HtrA2 contributes to uncontrolled cell death with Parkinsonian phenotype. Int J Biol Macromol. 2021;180:97-111.
-
Lin CH, Chen ML, Chen GS, Tai CH, Wu RM. Novel variant Pro143Ala in HTRA2 contributes to Parkinson’s disease by inducing hyperphosphorylation of HTRA2 protein in mitochondria. Hum Genet. 2011;130(6):817-827.
Tables
Table 1
Additional Information
Publisher’s Note
Bayrakol MP remains neutral with regard to jurisdictional and institutional claims.
Rights and Permissions
About This Article
How to Cite This Article
Duygu Kirkik, Abdurrahman Umut Tuyel, Sevgi Kalkanlı Taş, HTRA2 and its role in parkinson’s disease: Genetic and functional insights. Ann Clin Anal Med 2026;17(Suppl 2):S147-151. doi: 10.4328/ACAM.22741
- Received:
- May 15, 2025
- Accepted:
- June 23, 2025
- Published Online:
- July 7, 2025
- Printed:
- March 20, 2026

